In the first paper in this series, we asked a narrow question: if you stay inside a tightly comparable 16S/MaAsLin-style family of diet-microbiome studies, do published coefficient tables agree enough to be reused responsibly?
The answer was yes, with enough agreement to justify a reference layer.
That solved only the first problem.
Much of the most valuable newer microbiome literature no longer lives in that older 16S coefficient space. Large recent cohorts use shotgun metagenomics, centered-log-ratio (CLR) transformations, and different statistical workflows. They often contain broader phenotype coverage, better medication data, and more clinically structured variables. They also break naive comparability.
So the next question was obvious: can selected newer cohorts be brought into the same evidence program without pretending that all published coefficients are naturally interchangeable?
This post summarizes our second paper, which tackles that bridge problem.
Why the second paper was necessary
After the first paper, we had a validated 16S reference layer built from three cohorts:
- LifeLines-DEEP
- Flemish Gut Flora Project
- American Gut Project re-run into a comparable format
That reference layer was useful, but incomplete. It captured only a narrow slice of the public evidence base. Many of the larger and more modern cohorts now driving the field use different sequencing technologies and different transformed abundance spaces. Those studies often add exactly the kinds of variables a broader evidence layer needs: medication exposures, clinically structured host phenotypes, symptom profiles, and richer health-record-linked metadata.
In other words, the evidence base was expanding faster than the comparable part of the evidence base.
That is why paper 2 exists. It is not about claiming that older and newer cohorts are suddenly equivalent. It is about asking whether some of those newer cohorts can be connected back to the validated reference layer through an explicit calibration step.
The problem: mapped coverage is not the same as validated reuse
It is easy to overstate what harmonization means.
You can map factors across cohorts semantically and still have coefficient spaces that are not directly commensurable. A variable can line up by concept while the reported effect size lives in a different transform space. Once one cohort reports arcsine-square-root 16S coefficients and another reports CLR-based shotgun coefficients, raw magnitude comparison stops being trustworthy.
That means a broader harmonized panel creates two very different kinds of progress:
- mapped coverage, where more factors and taxa can be routed into a common framework
- validated overlap, where shared observations are strong enough to support actual cross-cohort comparison
Paper 2 is mainly about keeping those two ideas separate.
What we added
We expanded the panel from three cohorts to five by adding two shotgun/CLR-based studies:
- Dutch Microbiome Project (DMP) - 8,208 participants, broad phenotype coverage, large medication layer
- Estonian Microbiome Project (EstMB) - 2,509 participants, independent cohort with a different CLR-based workflow
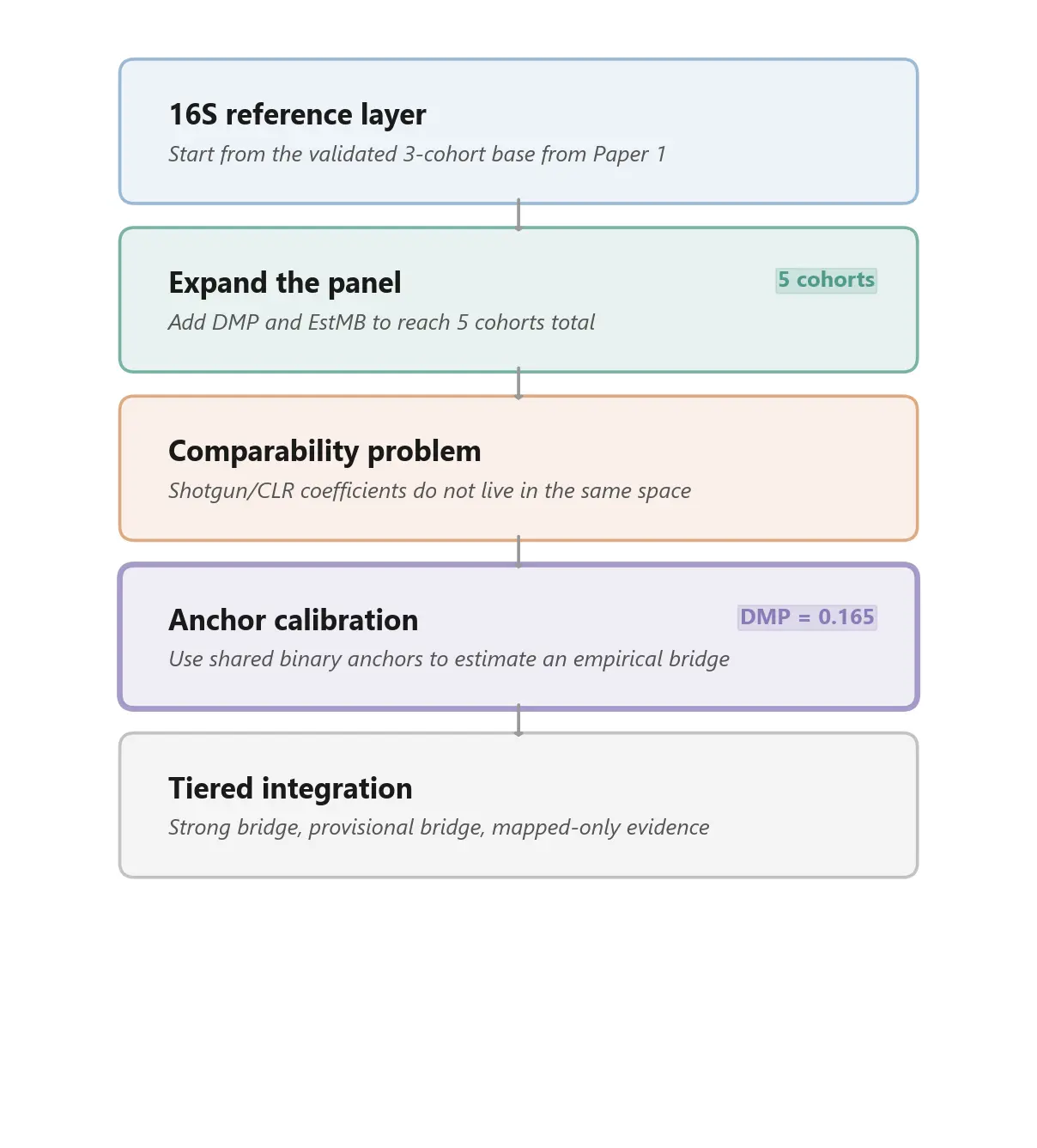

This expansion materially changed the evidence map. It especially widened coverage in:
- medications
- clinically structured exposures
- GI symptoms
- mental health variables
- early-life and health-record-linked factors
That is useful. But it also makes the comparability problem impossible to ignore. More mapped factors do not automatically mean more trustworthy coefficient reuse.
The bridge idea: anchor calibration
Our solution in this paper is what we call empirical anchor calibration.
The intuition is simple. If a newer target cohort and a validated reference cohort share some of the same factor-taxon associations, and those shared pairs agree on direction, then those overlaps can be used as anchors. The cleanest anchors are binary predictors, because they reduce the risk that a coefficient ratio is being driven by differences in predictor scaling rather than by the abundance transform itself.
So instead of treating CLR-based coefficients as if they lived on the same scale as the 16S reference layer, we asked whether shared binary anchors could estimate an empirical bridge back to that reference space.
That is a much narrower claim than universal comparability. It is also a much more defensible one.
What we found
The strongest bridge in the paper is DMP versus Zhernakova 2016.
This is the cleanest available cross-method comparison in the current panel because both cohorts come from the broader LifeLines setting. That shared provenance is a methodological caution, but it also makes this pair the best place to test whether a 16S reference cohort can be connected to a modern shotgun/CLR cohort without total collapse in agreement.
The result was encouraging:
- 100% sign agreement at species level across 12 shared significant pairs
- 74.2% sign agreement at genus level across 31 shared significant observations
- a positive genus-level rank correlation ($\rho$ = +0.416)
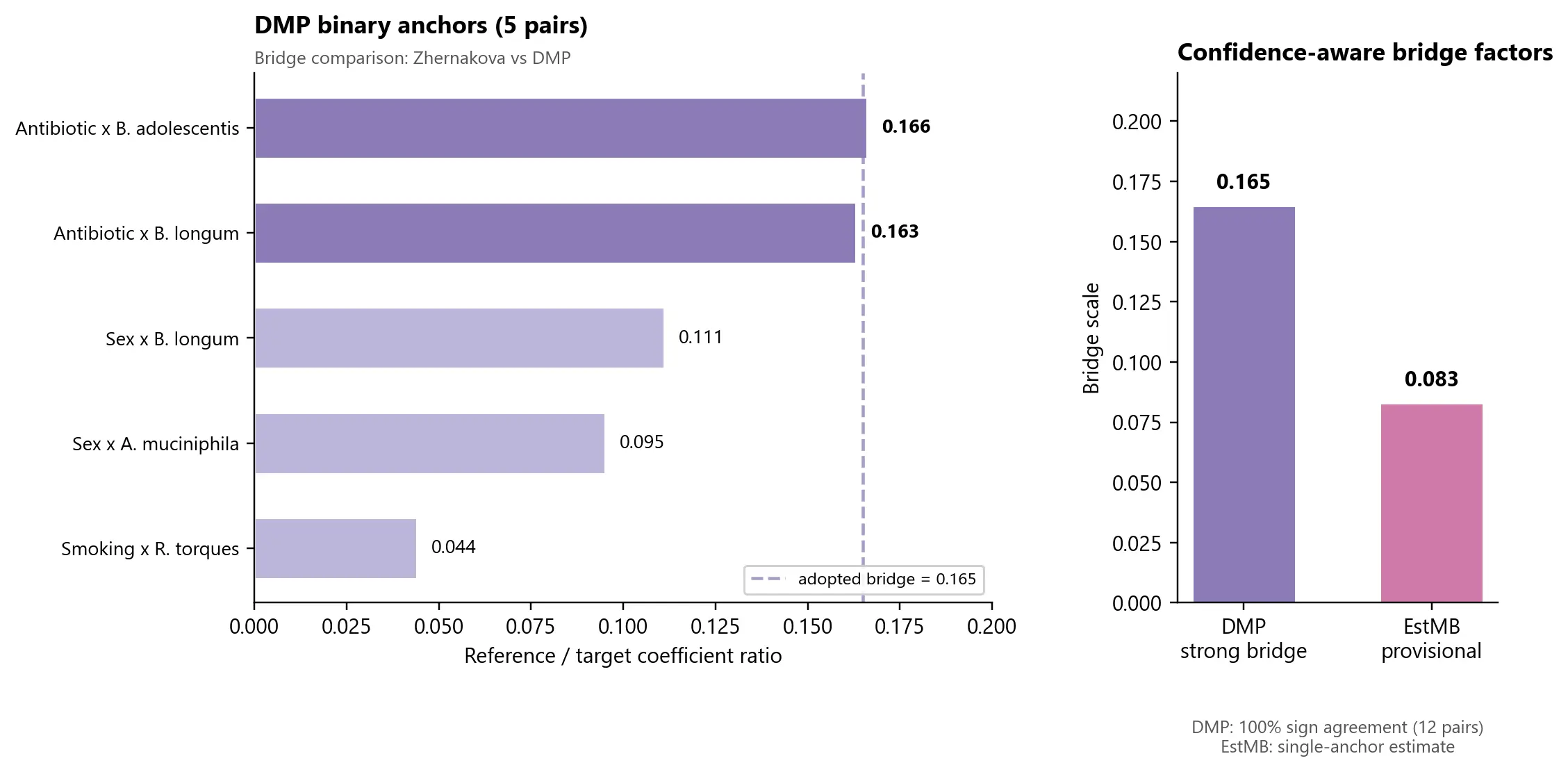
The binary-anchor subset within that bridge produced a tight high-confidence cluster around a scale factor of 0.165 for DMP relative to the 16S reference space.
That does not mean DMP is "the same" as the older reference cohorts. It means DMP looks bridgeable enough to enter the evidence layer through calibrated, explicitly qualified reuse.
Why this matters
This is the methodological step that turns a narrow validated reference layer into the beginning of a broader evidence program.
With DMP and EstMB, the mapped panel becomes much richer, especially in domains the earlier 16S cohorts barely covered. Medication-related signals are the clearest example. The expanded panel now contains substantially more structured evidence around proton pump inhibitors, metformin, statins, and related clinical exposures than the original three-cohort layer could support on its own.
But paper 2 is not a story about "more data, therefore better." It is a story about tiered inclusion.
Some cohorts can be bridged strongly. Some can only be carried forward provisionally. Some method families will remain outside this bridge entirely until a different framework exists.
That distinction matters if you want to build on the literature without overselling what has actually been validated.
The weaker bridge is still informative
The second new cohort, EstMB, matters for a different reason.
It is independent of LifeLines, so it tests whether the bridge idea survives outside the especially favorable DMP setting. Right now, the answer is only partially. EstMB yields a provisional scale estimate of 0.083, but that estimate currently rests on a single binary anchor pair.
So EstMB cannot be treated as a bridge of equal confidence to DMP.
That is not a failure. It is exactly the kind of result a staged evidence program should be willing to report. A thin bridge should be carried forward as thin, not promoted into false certainty.
One of the main lessons: expansion outruns validation
One of the most important outputs of paper 2 is conceptual rather than numerical.
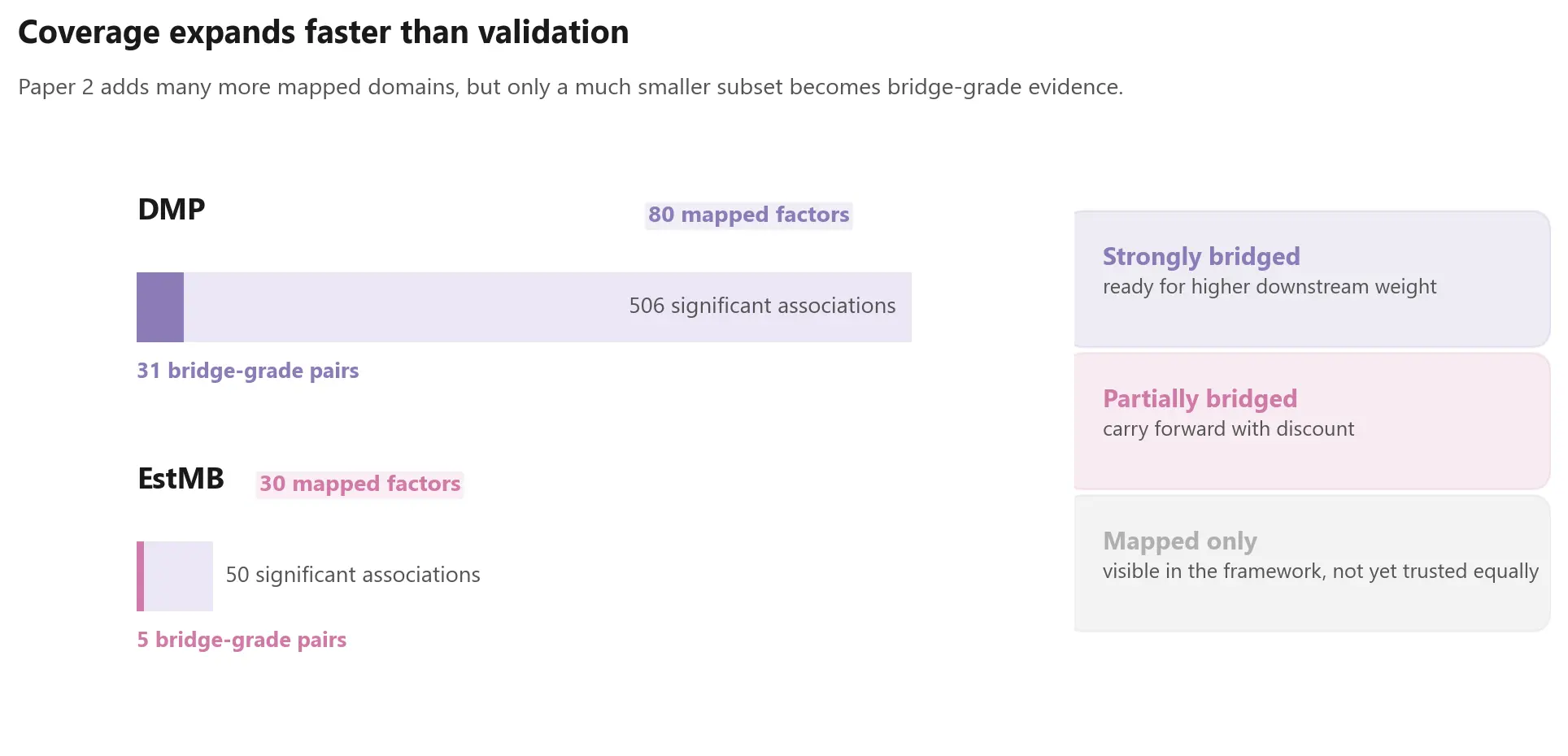
As the panel grows, mapped signal volume grows faster than directly validated overlap. DMP contributes hundreds of significant associations, but only a much smaller subset sits inside the doubly significant bridge space needed for calibration-grade comparison. EstMB is thinner still.
That asymmetry is not an annoyance around the edges of the analysis. It is the central fact that has to shape downstream model-building.
Any serious integration layer built from public microbiome results should distinguish between:
- strongly bridged evidence
- partially bridged evidence
- mapped-but-not-yet-validated evidence

What this means for Biome Bliss
For Biome Bliss, this paper is the second stage in a larger architecture.
Paper 1 established that coefficient reuse is possible inside a narrow compatible family. Paper 2 asks whether that reuse can be extended across an important method boundary.
The answer is encouraging, but conditional:
- some modern CLR-based cohorts appear bridgeable
- stronger bridges deserve more downstream weight
- weaker bridges should remain discounted
- and not every published method family belongs in the same integrated layer
That is the standard we want the project to follow. Not all evidence enters on equal terms. Each cohort earns its way in through comparability, validation, and explicit confidence handling.
What we are releasing
We are releasing the harmonization metadata, anchor-computation summaries, concordance results, and coverage metadata needed to inspect the logic of the bridge.
We are not releasing integrated model-ready coefficient tables or the downstream weighting stack built on top of them. Those remain proprietary parts of the broader Biome Bliss infrastructure.
Where this leaves the series
Taken together, the first two papers describe a more disciplined way to build on public microbiome evidence.
The first paper established a narrow validated reference layer. The second paper shows how selected newer cohorts can be connected back to that layer through empirical calibration rather than naive pooling.
That still leaves major parts of the literature outside the bridge. Some future cohorts may need different calibration logic entirely. But we think this is the right order of operations:
first validate, then bridge, then integrate, and only then trust the broader evidence layer enough to use it downstream.
Marvin Uhlmann and Maria Otworowska Biome Bliss Research, BlissLabs OÜ
Full technical paper: "Anchor calibration for extending diet-microbiome coefficient harmonization beyond 16S/MaAsLin cohorts."