If you want to build anything serious on top of gut microbiome research, there is an obvious question you have to answer first: when two large cohorts report associations between the same dietary factor and the same bacterial genus, do they actually agree?
That question matters for more than academic neatness. Investors, collaborators, and technically literate readers should be skeptical of any company claiming to turn microbiome papers into usable guidance unless it can show that the underlying evidence is at least directionally stable across independent cohorts.
This post summarizes the first validation step in our public evidence program at Biome Bliss Research. The goal was not to produce a consumer headline or a "gut health" story. The goal was narrower and more important: test whether published coefficient tables from comparable diet-microbiome studies are consistent enough to support disciplined downstream reuse.
This first step was deliberately narrow. Before asking whether newer shotgun and CLR-based cohorts can be brought into the same evidence layer, we wanted to establish that reuse works at all inside the most comparable published family.
The infrastructure problem behind microbiome products
The published diet-microbiome literature is now large enough that almost anyone can assemble a narrative from it. That is precisely the problem. Most studies use different questionnaires, different sequencing approaches, different statistical frameworks, and different variable encodings. Two papers may appear to report "the same" association while actually measuring different constructs in different units.
That means the real bottleneck is not the existence of published findings. It is coefficient portability. Can published results be compared, screened, and integrated without collapsing into a pile of nominally similar but methodologically incompatible signals?
Our view is simple: if you cannot show that comparable studies agree on the basics, you should not build a harmonized evidence layer from them.
What we tested
We did not try to compare the entire microbiome literature. That would have been methodologically unserious. Instead, we defined a narrow compatibility condition: 16S sequencing, MaAsLin-family regression, and factor encodings close enough to make coefficient-level comparison meaningful.
Out of eleven candidate cohorts, only three passed that filter.
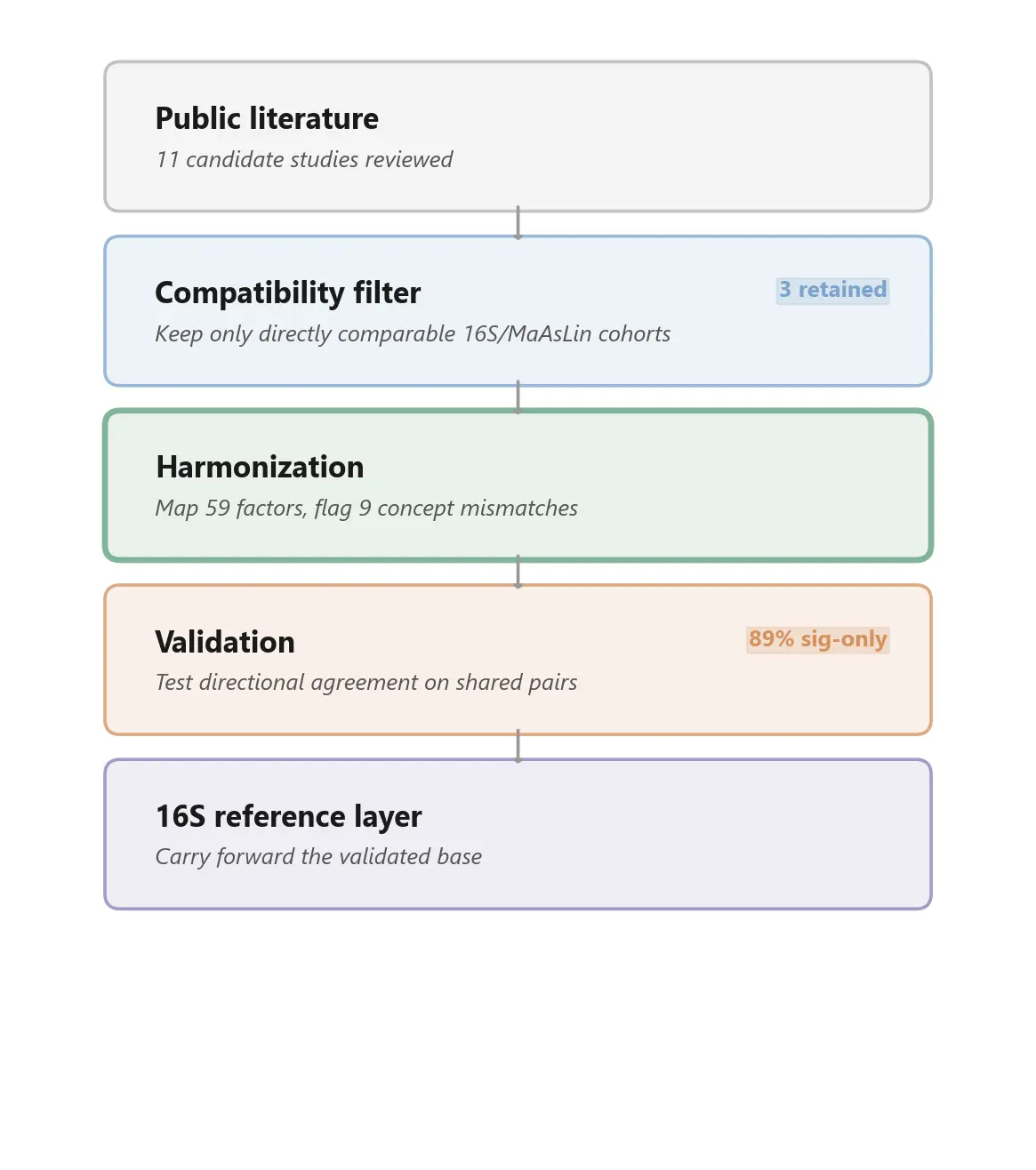

Those three were:
- LifeLines-DEEP - 1,135 Dutch adults, published in Science (2016)
- Flemish Gut Flora Project - 1,106 Belgian adults, also Science (2016)
- American Gut Project - 4,376 mostly American adults, which we re-analysed ourselves from publicly available data to make it compatible with the European studies
We mapped 59 dietary and lifestyle factors across the three cohorts and explicitly flagged nine cases where variables looked similar by name but differed in meaning or scale. That mismatch catalogue is not a side note. It is part of the point. Responsible reuse depends on identifying where naive keyword matching would create fake disagreement or fake agreement.
We then asked the simplest validation question available: when two cohorts both report a coefficient for the same factor and the same genus, do they agree on direction?
What we found
The answer was yes, within this compatible methodological family and within the limited overlap that currently exists.
Among the strongest shared signals - associations significant in both cohorts - 89% pointed the same way in the largest comparison.
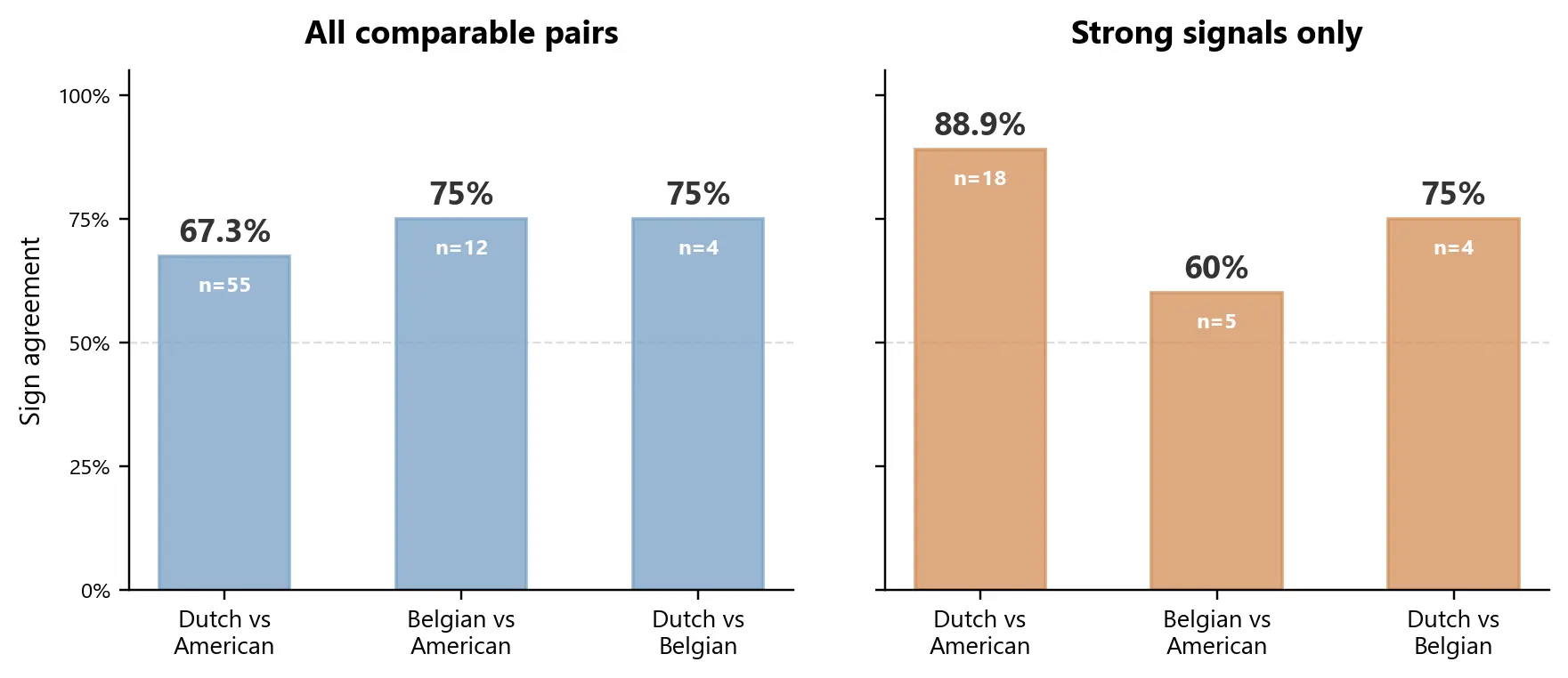
Specifically, 16 of 18 shared significant pairs agreed between the Dutch LifeLines-DEEP cohort and the re-run American Gut cohort. The coefficient magnitudes were also positively correlated, which suggests that this was not only a direction-of-effect story.
When we widened the comparison to include all comparable pairs regardless of statistical strength, agreement fell to 67%. That is not surprising. Weak signals are noisier, and any serious integration framework should distinguish between broad overlap and high-confidence overlap.
Only two associations were testable across all three cohorts, both involving Bristol stool type:
- Looser stools = less Bacteroides - agreed in all three cohorts
- Looser stools = more Roseburia - agreed in all three cohorts
These are not flashy findings, but they are useful ones. They match known transit-time physiology and serve as clean biological anchors inside a sparse overlap set.
What this means
This result does not mean microbiome science is settled, nor does it mean published coefficient tables are interchangeable in raw form. The overlap here is limited. Three Western cohorts are a narrow slice of global dietary diversity, and only 17 of the 59 harmonized factors were comparable across at least two studies.
What it does mean is that careful compatibility screening, concept-level harmonization, and explicit mismatch handling can recover a shared comparison space where agreement is strong enough to justify further work.
For researchers, that is a useful methodological result. For collaborators and investors, it is evidence that we are not treating "the literature" as an undifferentiated blob. We are building an evidence layer in stages, starting with the subset of public results that can first earn trust through validation.
In our case, that means establishing a reference layer before attempting harder bridge problems. The next methodological question is not whether more cohorts exist, but whether cohorts generated with different sequencing and transformation pipelines can be added without breaking comparability.
Why this matters for Biome Bliss
Biome Bliss is building a dietary guidance tool on top of public and structured evidence about how food patterns relate to gut microbiome composition. That only becomes defensible if the evidence pipeline is more rigorous than "collect a lot of papers and average them."
This validation exercise is the first layer of that pipeline. Before integrating cohorts into downstream models, we want to know:
- which studies are comparable at all
- where agreement is real
- where disagreement is probably methodological
- and where the overlap is still too thin to justify strong claims
That is the standard we think this space needs more of.
What we are releasing
We are releasing the validation logic, the harmonization metadata, and the summary concordance statistics that allow others to inspect the reasoning.
We are not releasing integrated model-ready weights or the full downstream modelling stack. Those are part of the proprietary infrastructure we are building on top of the validated public evidence base.
What comes next
This is the first paper in a staged series. It establishes the narrow 16S/MaAsLin reference layer that later extensions have to bridge back to. Additional cohorts can be brought into the framework only after they pass the same basic test: methodological compatibility first, harmonization second, validation before integration.
That is slower than simply pooling everything that looks relevant. We think it is also the only credible way to build on public microbiome evidence without overstating what the literature can support.
The next step in the series takes exactly that harder case: whether selected newer shotgun/CLR cohorts can be connected back to this reference layer through empirical anchor calibration, expanding coverage without pretending that all coefficient spaces are naturally interchangeable.
Marvin Uhlmann and Maria Otworowska Biome Bliss Research, BlissLabs OÜ
Full technical paper: "Building a harmonized public evidence base for diet-microbiome modelling: initial validation across three 16S cohorts."