The first two papers in this series solved one kind of problem. Paper 1 showed that published 16S coefficient tables can agree within a narrow compatible family. Paper 2 showed that selected newer shotgun/CLR cohorts can be bridged back to that reference layer through empirical anchor calibration.
Both papers operated within a shared assumption: each cohort reports associations between individual dietary factors and specific bacterial taxa. That made cross-cohort comparison methodologically tractable, even when the statistical transforms and sequencing technologies differed.
The Human Phenotype Project breaks that assumption.
Why HPP matters and why it is difficult
HPP (Segev et al. 2026) is, by several measures, the single most important public diet-microbiome resource currently available. It combines 10,064 participants with shotgun metagenomics, 669 species, detailed app-based dietary logging, and longitudinal follow-up. In scale and depth, it materially exceeds most cohorts in our existing five-cohort panel.
The problem is that HPP's published supplementary outputs emphasize species associations with composite diet quality scores - not with individual foods. That is a scientifically reasonable design choice within a single study. But for a food-factor harmonization framework, it creates a structural mismatch that cannot be solved by the same calibration logic used in Paper 2.
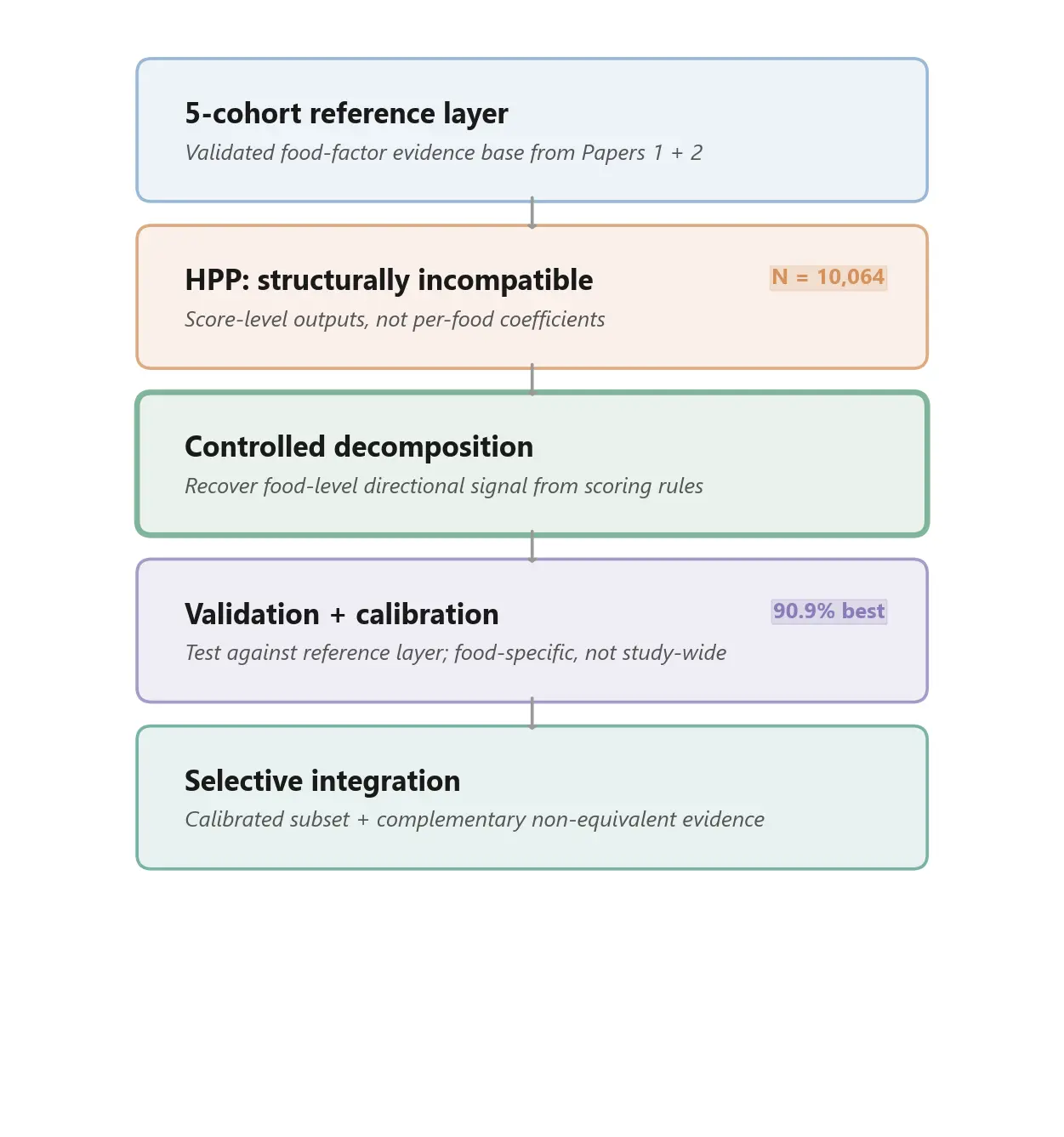

This is not a quality problem. It is a format problem. And it is the kind of problem that will become more common, not less, as the public microbiome literature matures.
What we tried: recovering food-level signal from score-level associations
HPP publishes scoring rules for its diet quality indices. Those scoring rules specify which foods contribute to which scores, and in what direction. If a diet score has a strong positive association with a particular species, and a specific food contributes positively to that score, then the food-species direction can be inferred.
That inference is bounded. It cannot recover exact coefficients. It cannot separate confounded food contributions within a single score. But it can recover directional evidence - enough to test whether HPP's published outputs are broadly consistent with independently derived food-taxon associations from the existing five-cohort reference layer.
What we found
The recovered HPP food-level signals showed substantial directional agreement with the reference framework:
- 73.3% agreement across the full testable overlap
- 83.3% after excluding known population-specific confounds (Israeli dairy/plant structure)
- 90.9% after also excluding a near-zero ambiguous case
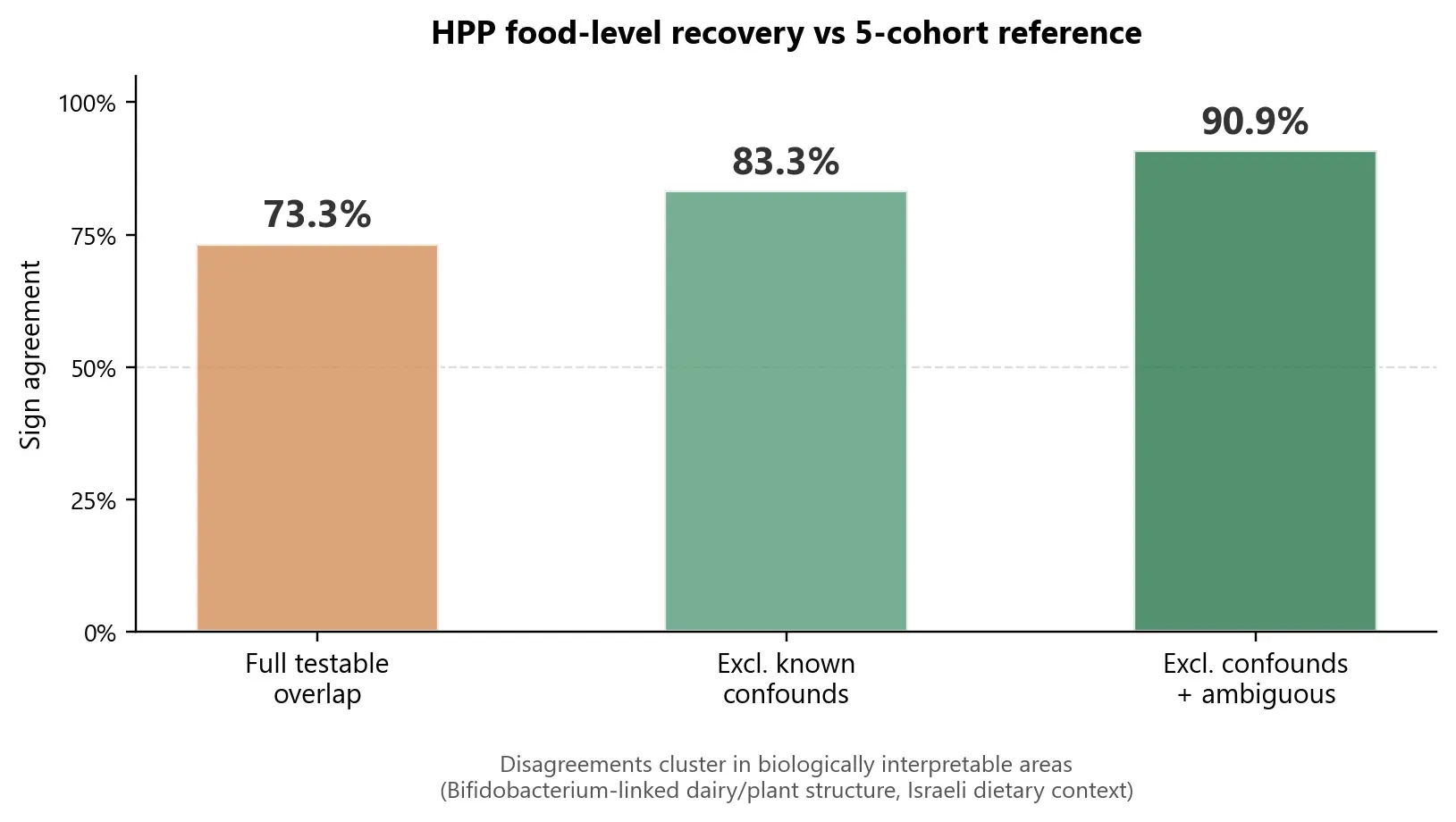
The disagreements were not random. They clustered around biologically interpretable areas - especially Bifidobacterium-linked dairy and plant structure, where Israeli dietary context plausibly shifts food-taxon associations relative to European and American cohorts.
That matters. Patterned, interpretable disagreement is more informative than uniform noise. It suggests the decomposition is recovering real cohort-specific signal, not manufacturing agreement.
The calibration problem is food-specific, not study-wide
After confirming directional validity, we tested whether the recovered HPP food-level values could be placed onto the existing reference scale. The answer: not with a single study-wide conversion constant. Cross-food heterogeneity was too large to support that claim.
But a narrower result did hold. For a validated subset of food-level signals, calibration behaved consistently enough to support bounded, food-specific reuse within the harmonized framework.
This is the central finding of the paper: HPP can be partially bridged, but only selectively and only under explicit food-level validation.
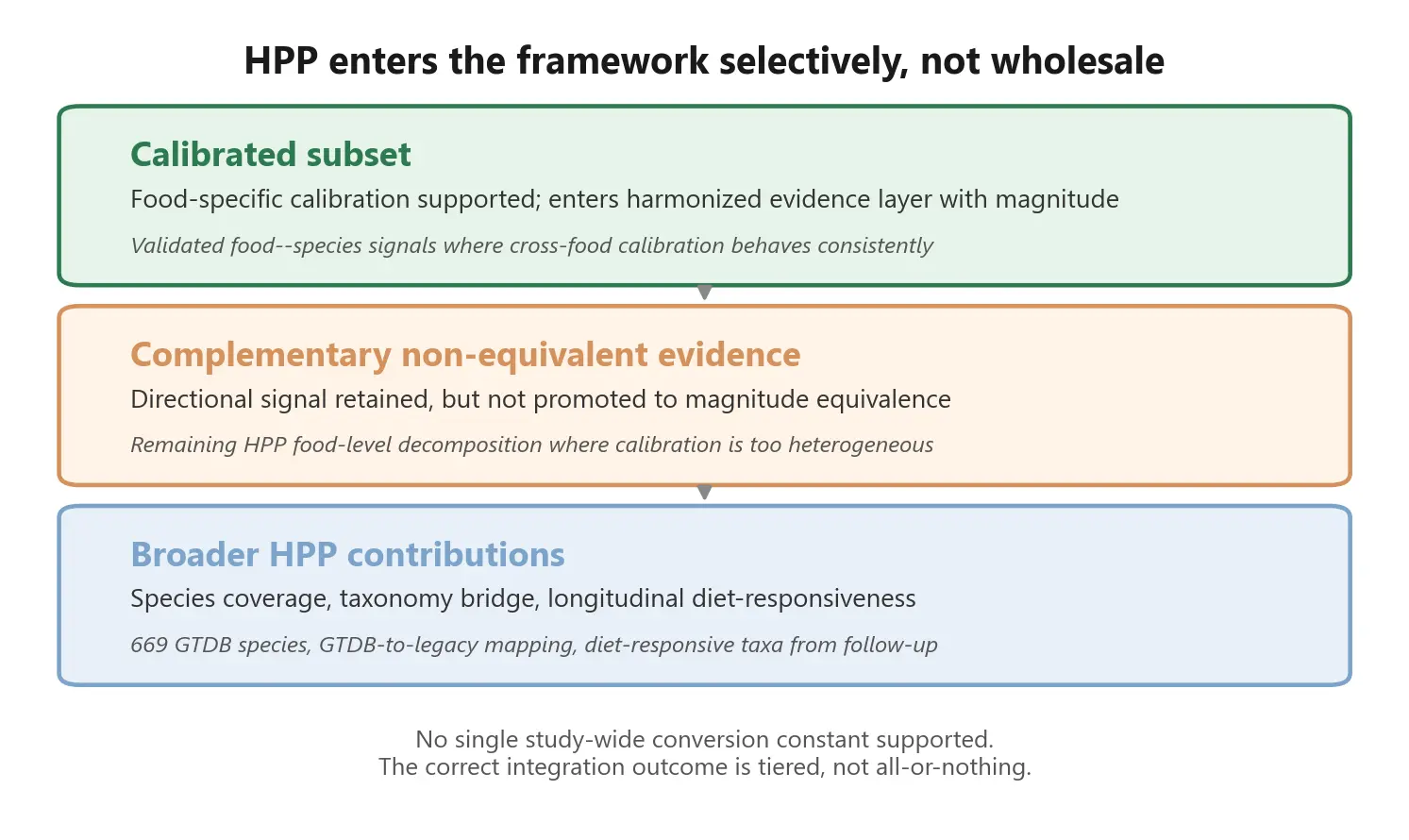

Why selective integration is the right answer
It would be tempting to force HPP into the framework wholesale. The data volume is large, the species coverage is excellent, and the longitudinal validation is unique. But doing so would overstate what has actually been validated.
The correct outcome for a responsible evidence program is often partial inclusion rather than all-or-nothing. A calibrated subset enters the harmonized evidence layer with magnitude information. The remainder is retained as complementary, non-equivalent evidence - visible in the framework, useful for context and coverage expansion, but not promoted to full coefficient equivalence.
That tiered structure is not a weakness. It is the discipline that makes the broader evidence layer trustworthy.
What HPP adds beyond the calibrated subset
Even outside the directly calibrated subset, HPP strengthens the framework in three important ways:
- Broader species coverage - 669 GTDB-level species, far more than the 16S-era cohorts
- Taxonomy bridge - GTDB-to-legacy name mapping that makes a larger modern species space interpretable within the earlier framework
- Longitudinal diet-responsiveness - species that respond to dietary change over follow-up, a dimension largely absent from the earlier cohort panel
These contributions matter even where direct magnitude-level integration is not justified.
Where this leaves the series
The three papers now form a clear progression:
Paper 1 established that published food/factor coefficients can agree within a narrow directly comparable family.
Paper 2 showed that selected cross-method bridges are possible when calibration is used carefully.
Paper 3 extends that logic to a fundamentally different form of incompatibility - not just a different transform, but a different unit of published dietary evidence altogether.
Together, the series is a statement about how public microbiome evidence should be integrated: carefully, with explicit compatibility rules, with validation before inclusion, and with the willingness to leave some evidence only partially absorbed rather than forcing false equivalence.
What we are releasing
We are releasing the validation logic, food-level recovery summaries, calibration structure documentation, and taxonomy bridge metadata.
We are not releasing integrated model-ready coefficient tables or downstream weighting logic. Those remain part of the proprietary Biome Bliss infrastructure.
Marvin Uhlmann and Maria Otworowska Biome Bliss Research, BlissLabs OÜ
Full technical paper: "Integrating diet-score microbiome evidence into a harmonized multi-cohort framework: controlled decomposition and calibration of the Human Phenotype Project."